Research
Multi-omics-guided genome synthesis and design
Rational genome design and genetic code engineering enable three key innovations: (i) broad resistance to natural viruses, (ii) prevention of genetic information flow between engineered organisms and natural species, and (iii) the capability to biosynthesize entirely new classes of genetically encoded polymers.
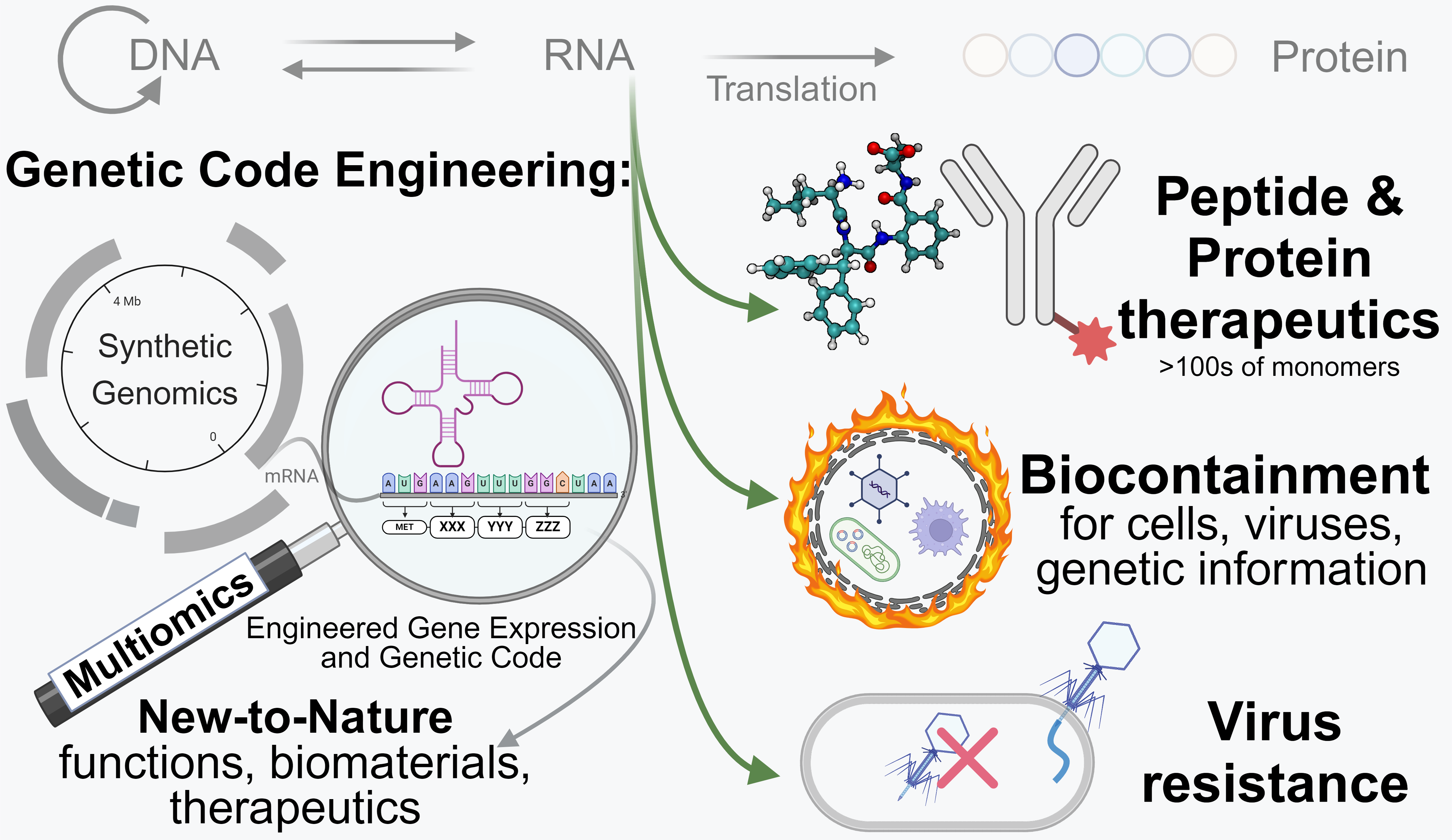
Multi-omics-guided synthesis and rescue of radically recoded E. coli genomes.
We have developed a multi-omics-guided directed evolution and genome-design strategy that rapidly debugs synthetic chromosomal regions and restores fitness, paving the way for the synthesis of functional, high-fitness genomes with radical changes. This technology serves as foundation for our current research projects.
Amino-acid-swapped genetic code providing virus-resistance and biocontainment.
Illustration by Behnoush Hajian; celline.design
Genetic firewall for virus-resistance and biocontainment
The universal nature of the genetic code allows organisms to exchange functions through horizontal gene transfer (HGT) and enables recombinant gene expression in heterologous hosts. However, the shared language of the same code permits the undesired spread of antibiotic-, herbicide-, and pesticide-resistance genes and allows viruses to cause diseases.
We have developed a technology that renders cells resistant to natural viruses and biocontains cells and their genetic information by establishing a genetic-code-based firewall.
This genetic-code-based firewall renders Escherichia coli cells resistant to viruses (including bacteriophages in environmental samples) by mistranslating viral proteomes and prevents the escape of synthetic genetic information. Simultaneously, we biocontain this virus-resistant host through dependence on an amino acid not found in nature. This work establishes a strategy to make organisms safely resistant to natural viruses and prevent genetic information flow into and out of Genetically Modified Organisms (GMOs).
Highlighted in Nature Biotechnology; New Scientist; Synthetic Biology; Science; U.S. Department of Energy; Nature News & Views.
Interviews in Nature Podcast, Drug Discovery News, The Scientist, and Harvard Medical School News.
Accelerated directed evolution in diverse bacteria
We developed a method that enables the precise mutagenesis of multiple, long genomic segments in multiple species without off-target modifications. This technology (DIvERGE) enables the exploration of vast numbers of combinatorial genetic alterations in their native genomic context and allows accelerated directed evolution.
We demonstrated that our broad-host-range genome engineering (pORTMAGE; Nyerges, A. et al., PNAS, 2016, PMID: 26884157) enables the translation of chemical DNA-synthesis-based mutagenesis into focused genome-diversification with up to >1,000,000x the wild-type mutation rate.
DIvERGE + pORTMAGE enable precise, portable, multi-species directed evolution.
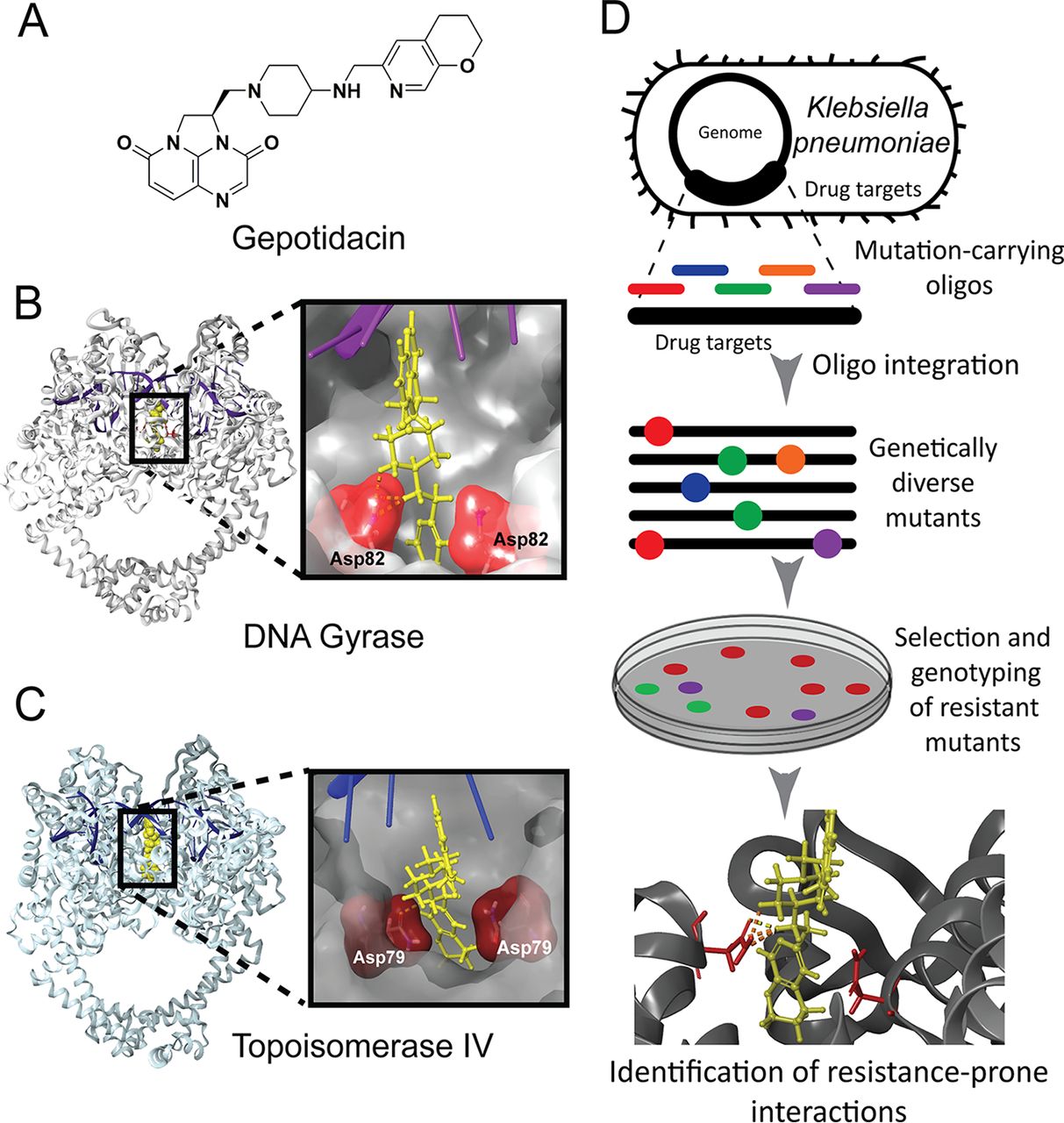
Using the combinatorial mutagenes of drug targets with DIvERGE, we identified previously undetected resistance processes for multiple antibiotics, including a clinical-stage antibiotic, gepotidacin (blujepa). Strikingly, gepotidacin's clinical trial later revealed the same resistance process in patients.
This technology was later outlicensed for drug development.
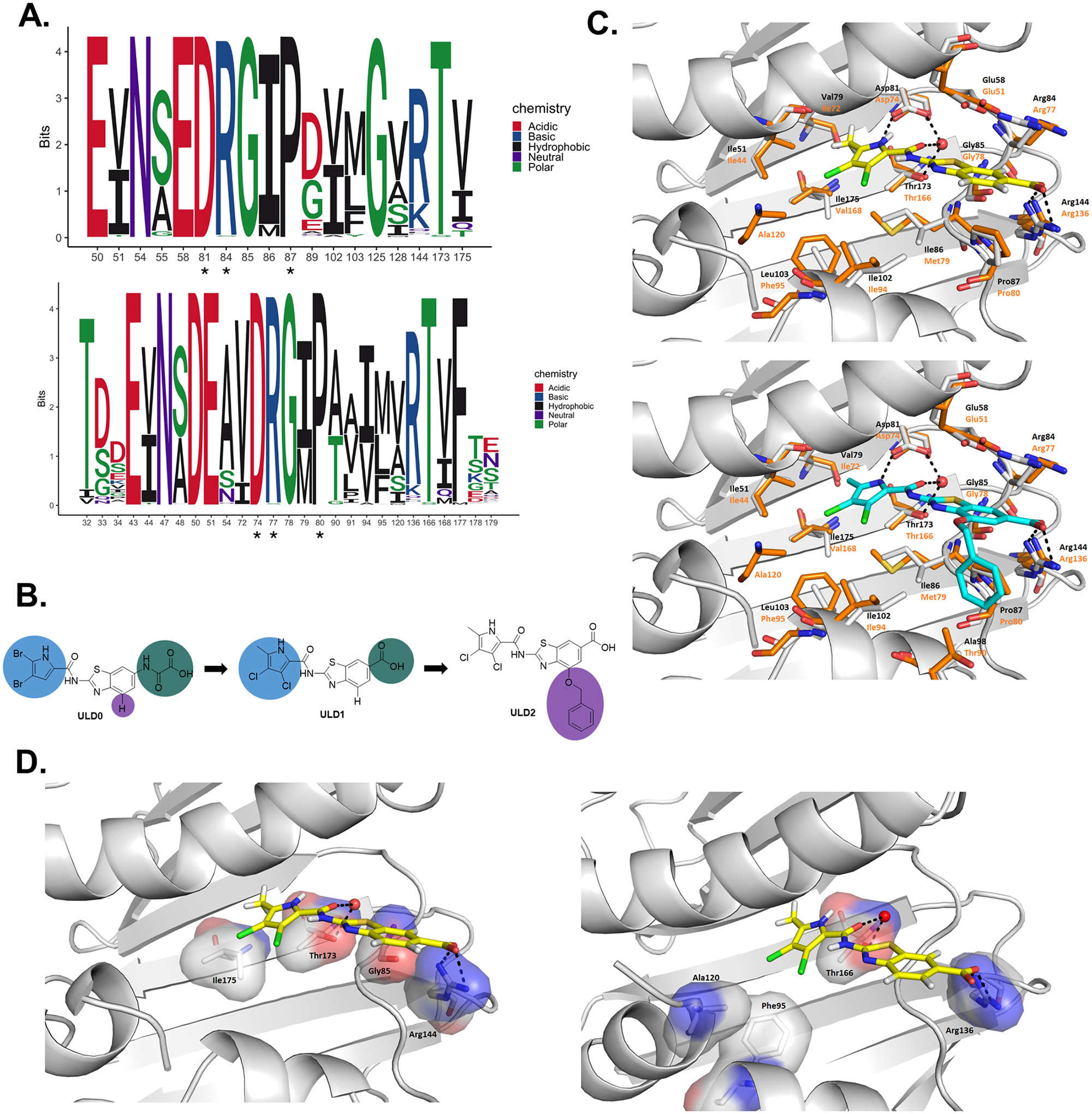
Rational design of dual-targeting DNA gyrase and topoisomerase IV inhibitors with suppressed resistance evolution.
Rational design of antibiotic drugs with limited resistance
In collaboration with Lucija Peterlin Masic's team — using structure-guided rational drug design, we developed a series of novel DNA gyrase and topoisomerase IV dual-targeting antibiotics.
We combined rational, target-based drug development with evolutionary analysis and the high-throughput prediction of resistance processes to identify key residues in drug-target interaction and suppress the evolution of drug resistance by rationally modifying our drug candidate.
This novel antibiotic displays broad activity against both drug-susceptible and multidrug-resistant Gram-positive bacterial pathogens, lacks toxicity in preclinical tests, was well-tolerated in mice, and demonstrated exceptional potency in mouse infection models.
See complete publication list →
Funding
Our research is supported by the U.S. National Institutes of Health and the U.S. Department of Energy.